There is a technology that turns materials, such as batteries, sensors, and new drugs into viruses, called bacteriophages. It is used as a platform to easily manipulate the genes of bacteriophages and make the desired protein molecules in bulk. It takes advantage of the virus’s rapid multiplication, and quantitative data is coming to the attention that will improve the way it relies only on past experience.
A joint research team, led by Professor Dong Woog Lee and Professor Jungki Ryu in the School of Energy and Chemical Engineering at UNIST, has presented ways to improve the platforms, which can be utilized as a database of the bacteriophage interactions. The study quantitatively measures the interaction between bacteriophages and substances on their surfaces.
The researchers also found that the dispersibility of materials that combine bacteriophages and carbon nanotubes (SWCNTs) can be controlled by pH. This result can be utilized in the production of carbon nanomaterials using bacteriophages.
Bacteriophages use bacteria such as Escherichia coli as a host to rapidly reproduce and mass reproduce their genetic material. It is also harmless to humans because it has the property of attacking only the host bacteria. A lot of research is underway to use these features to produce proteins for antibodies and new drugs. In addition, attempts are being made to apply to the nanomaterial field in that the size is very small at the nanometer level. It is intended to serve as a platform for producing or attaching desired protein molecules to the surface of bacteriophages.
Bacteriophage, which has a simple structure with genetic material and a protein shell surrounding it, is easy to manipulate. This allows them to create specific protein molecules on the surface, or to bind to specific materials. However, the lack of understanding of the forces acting between the surface protein molecules of the bacteriophage and other molecules has limited the ability to accurately design the desired material.
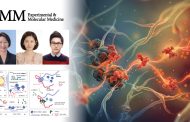
Experimental setup and contact angle measurements.
The researchers measured the pushing and pulling force between one of the bacteriophages, the M13 bacteriophage and the 5 other molecular groups, while changing the pH level. It used a device called SFA to measure directly at the molecular level, even at very close forces.
As a result, the adhesion energy between the phenyl group and the bacteriophage was the highest among the five molecular functional groups. This was due to the pulling force between the cyclic carbon structures of the protein and the phenyl group on the surface of the bacteriophage and the sticking of the hydrophobic substances in the aqueous solution.
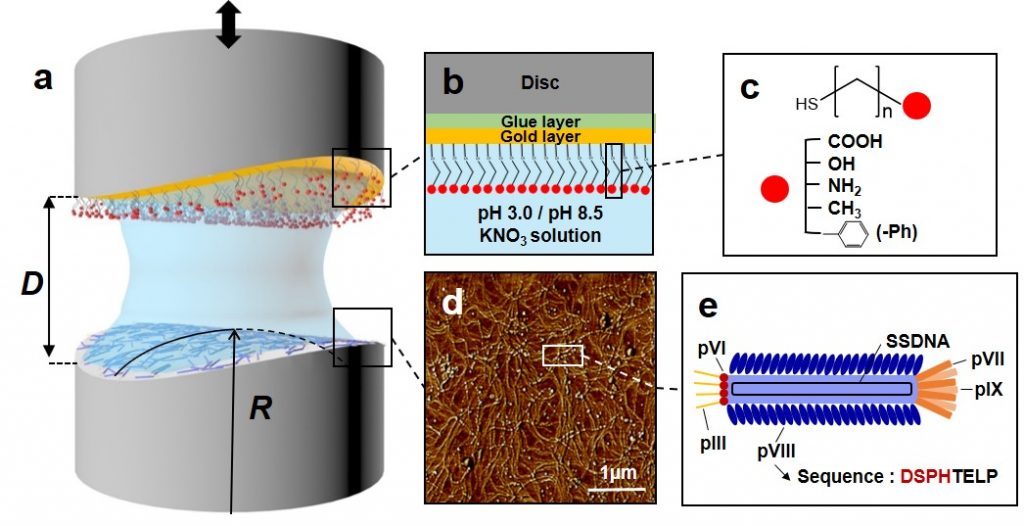
Based on the results, the researchers created a composite between the M13 bacteriophage and single-walled carbon nanotubes (SWCNTs). Afterwards, the bacteriophage adhered more strongly to carbon nanotubes under acidic conditions. This is because the pulling force of the annular carbon structure between each other and the pulling force of the two hydrophobic materials are strong.
The force to aggregate and disperse the bacteriophage complex was controlled by pH change. Under acidic conditions, proteins outside the bacteriophage bind more positively to the negatively charged carbon nanotubes. Because of this, the complexes were also stuck to each other, and tangled together.
On the other hand, under basic conditions, the bacteriophage external protein is negatively charged. In this case, the composites do not entangle because they repel each other with carbon nanotubes with the same negative charge. If this is applied, the pH can be controlled to easily control the density of the material attached to the bacteriophage.
“We believe that this study can provide a versatile platform to characterize various specific biomolecular interactions and enable better understanding and utilization of biomolecules,” says Professor Lee.
The findings of this research have been published in the journal, Communications Chemistry on August 20, 2019. This study has been supported by the National Research Foundation of Korea (NRF).
Journal Reference
Chanoong Lim et al., “Probing molecular mechanisms of M13 bacteriophage adhesion,” Communications Chemistry, (2019).